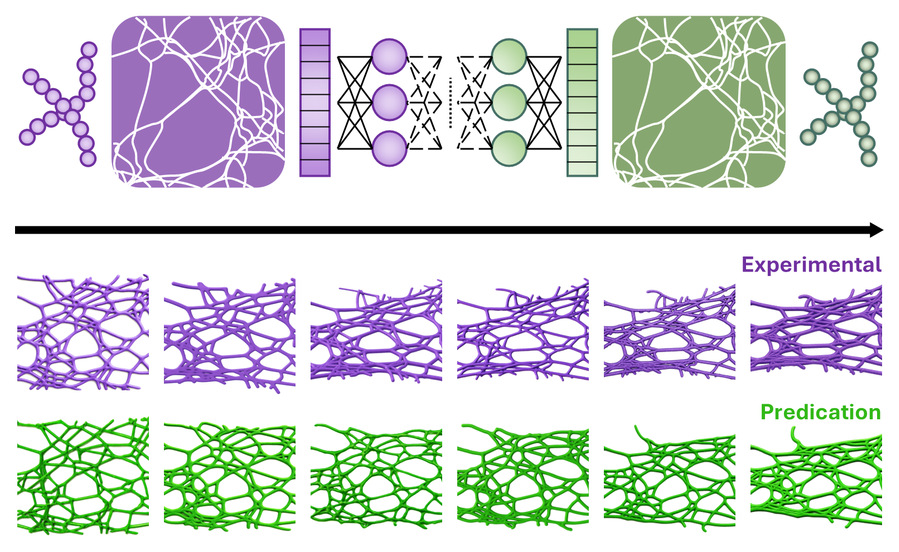
Disordered fibrous networks play vital mechanical roles but are difficult to model due to sparse connectivity and complex non-affine deformation. We introduce a physics-informed, graph-learning-based Network Mechanics Prediction (GNMP) that predicts deformation and stress–strain responses of 2D semiflexible networks with reliable accuracy and >10× efficiency gains over molecular dynamics. GNMP integrates graph-attention message passing, multi-scale physical embeddings, and a bond-length-guided scheduler to capture key non-affine rearrangements. Experiments on Poisson-ratio-aligned 3D-printed networks show consistent deformation motifs, descriptor trends, and comparable log-area changes (~0.46 vs ~0.24), while reducing geometry-inference time to ~0.13 s per step. GNMP offers a generalizable route for rapid, topology-aware mechanical prediction in fibrous network materials and supports accelerated design of biomimetic soft tissues, flexible conductors, and related network-based systems.
https://pubs.acs.org/doi/10.1021/acs.nanolett.5c04895